Research subject
- Structural and mechanistic basis of protein quality control systems in cells
Ongoing projects
- Determination of high-resolution structures of molecular chaperones and disulfide bond formation/cleavage enzymes involved in protein quality control
- Elucidation of the cellular mechanisms underlying the maintenance of protein and redox homeostasis by cell biological approaches
- Extensive analysis of the cellular redox network by proteomic approaches
Outline of our research
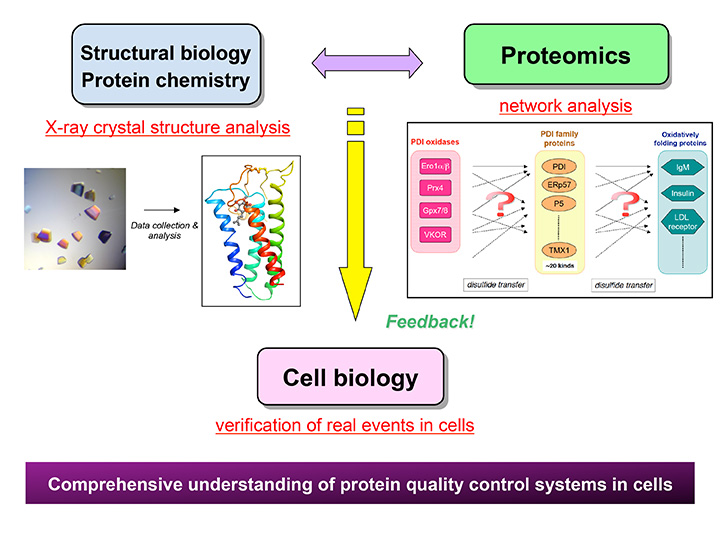
The biological kingdoms have evolved elaborate systems to assist proper protein folding in the cell. The formation and cleavage of disulfide bonds play important roles in maturation, assembly and functional regulation of proteins. Employing structural, biochemical, proteomic and cell biological approaches, we aim at the deep understanding of how the disulfide bond formation and cleavage systems cooperate to ensure the protein quality control in living cells. Structural and mechanistic insights gained in this work will reveal molecular mechanisms underlying the maintenance of protein and redox homeostasis and shed light on neurodegenerative diseases caused by abnormal proteins.
Selected Publications
- Inoue, M., Sakuta, N., Watanabe, S., Zhang, Y., Yoshikaie, K., Tanaka, Y., Ushioda, R., Kato, Y., Takagi, J., Tsukazaki, T., Nagata, K. and *Inaba, K. Structural basis of sarco/endoplasmic reticulum Ca2+-ATPase 2b regulation via transmembrane helix interplay Cell Reports, 27, 1221-1230 (2019)
- Okumura, M., Noi, K., Kanemura, S., Kinoshita, M., Saio, T., Inoue, Y., Hikima, T., Akiyama, S., Ogura, T. and *Inaba, K. Dynamic assembly of protein disulfide isomerase in catalysis of oxidative folding, Nat. Chem. Biol. 15, 499-509 (2019)
- Watanabe, S., Amagai, Y., Sannino, S., Tempio, T., Anelli, T., Harayama, M., Masui, S., Sorrentino, I., Yamada, M., Sitia, R. and *Inaba, K. Zinc regulates ERp44-dependent protein quality control in the early secretory pathway, Nat. Commun., 10, 603 (2019)
- Watanabe, S., Harayama, M., Kanemura, S., Sitia, R. and *Inaba, K. Structural basis of pH-dependent client binding by ERp44, a key regulator of protein secretion at the ER-Golgi interface, Proc. Natl. Acad. Sci. U S A. 114, 3224-3232 (2017)
- Arai, K., Takei, T., Okumura, M., Watanabe, S., Amagai, Y., Asahina, Y., Moroder, L., *Hojo, H., *Inaba, K. and *Iwaoka, M. “Preparation of selenoinsulin as a long-lasting insulin analog” Angewandte Chemie 56, 5522-5526 (2017)
- Vavassori, S., Cortini, M., Masui, S., Sannino, S., Anelli, T., Caserta, I., Fagioli, C., Mossuto, M., Fornili, A., van Anken, E., Degano, M., Inaba, K. and Sitia, R. “A pH-Regulated Quality Control Cycle for Surveillance of Secretory Protein Assembly” Mol. Cell 50, 783-792 (2013)
- Hagiwara, M., Maegawa, K., Suzuki, M., Ushioda, R., Araki, K., Matsimoto, Y., Hoseki, J., Nagata, K. and *Inaba, K. Structural basis of an ERAD pathway mediated by the ER-resident disulfide reductasse ERdj5. Mol. Cell 41, 432-444 (2011)
- Inaba, K.*, Masui, S., Iida, H. Vavassori, S., Sitia, R. and Suzuki, M. “Crystal structures of human Ero1α reveal the mechanisms of regulated and targeted oxidation of PDI” EMBO J 29, 3330-3343 (2010)
- Inaba, K.*, Murakami, S., Nakagawa, A., Iida, H., Kinjo, M., Ito, K. and Suzuki, M. “Dynamic nature of disulfide bond formation catalysts revealed by crystal structures of DsbB” EMBO J 28, 779-791 (2009)
- *Inaba, K., Murakami, S., Suzuki, M., Nakagawa, A., Yamashita, E., Okada, K. and *Ito, K. Crystal structure of the DsbB-DsbA complex reveals a mechanism of disulfide bond generation. Cell 127, 789-801 (2006)
Contact Information
“Please delete “/” after “@” in a mail address.”
Kenji Inaba
(TEL: 022-217-5604, E-mail: kinaba@/tagen.tohoku.ac.jp)
Satoshi Watanabe
(TEL: 022-217-5605, E-mail: stwata@/tagen.tohoku.ac.jp)